

| (10) Vanouse, preliminary research at SymbioticA, Perth. | (11) ibid. |
| Latent Figure Protocol, Paul Vanouse, 2007-09 | Vanouse projects |


| (10) Vanouse, preliminary research at SymbioticA, Perth. | (11) ibid. |
| The
“wet-biological” techniques used in the Latent
Figure
Protocol were researched throughout 2006 and are based in restriction
digestion of DNA samples and gel electrophoresis. The LFP
imaging
process relies upon knowing what size DNA is required for each band to
move at the proper speed to make the correct image. This is
essentially doing molecular biology IN REVERSE. Usually
scientists use imaging techniques to determine an organism’s
genetic sequence, whereas LFP utilizes known sequences in online
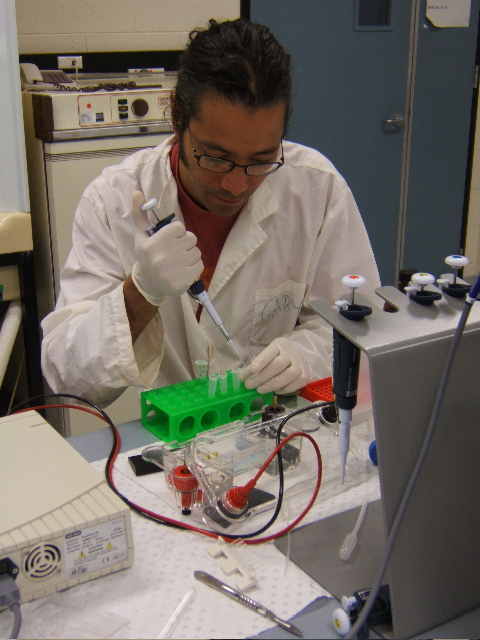
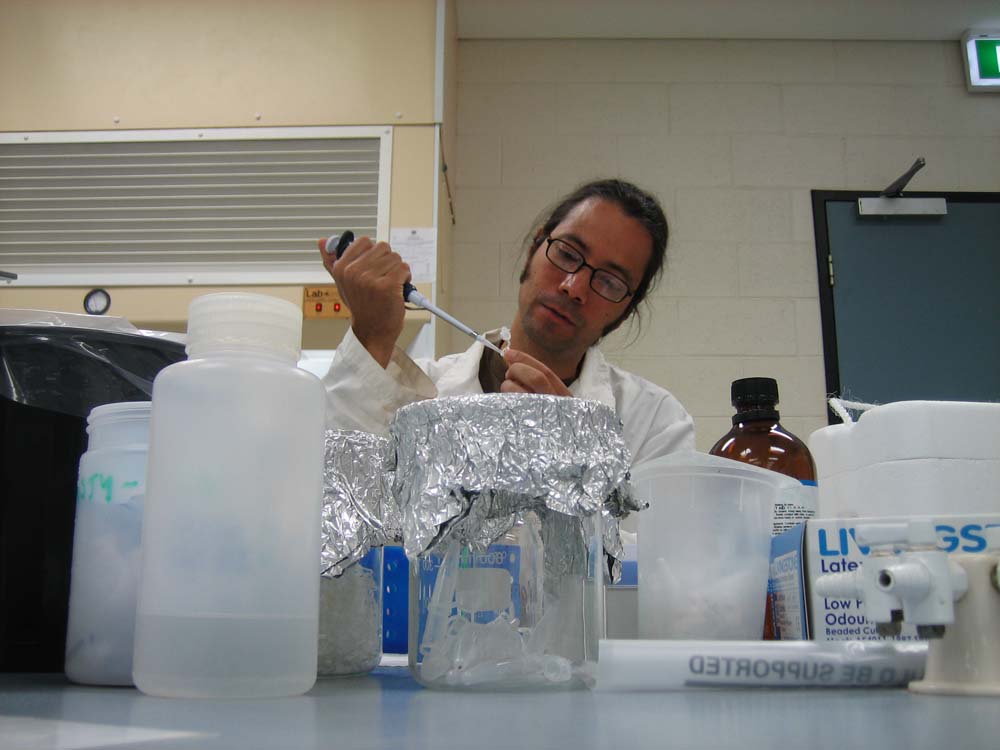
databases to produce “planned” images. These wet processes are consistent with my practice in emerging media forms. While I am deeply fascinated by many of these varied techno-scientific disciplines, I am interested in creatively “hacking” them with the aim of forcing the arcane codes of scientific meaning “to speak” in a broader cultural language. |
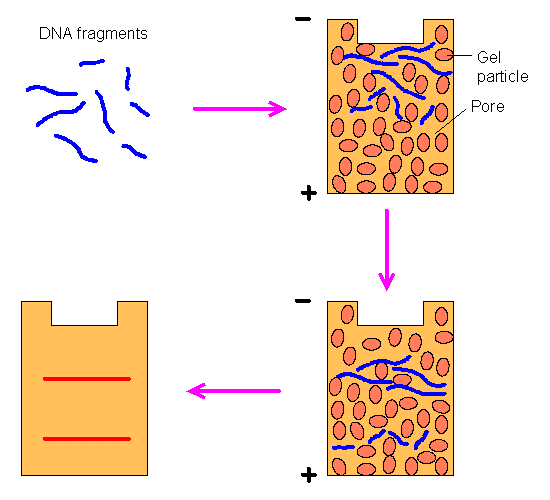
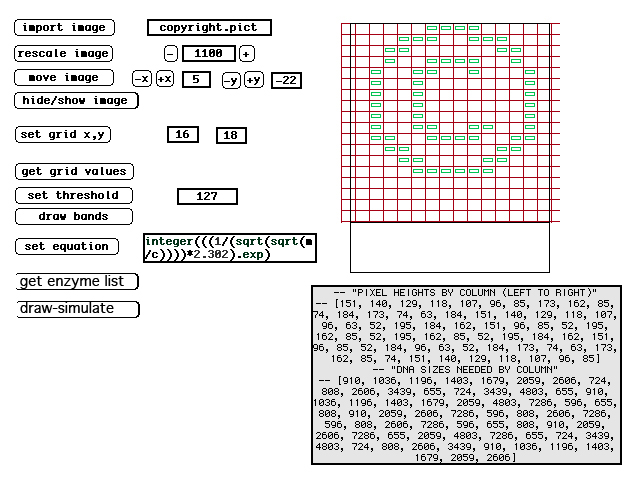
| (12) DNA moves through the gelatin at a rate inversely proportional to its size--thus smaller DNA moves faster. Note, this illustration shows the DNA moving downward, whereas LFP images show DNA moving upward. LFP utilizes horizontal gels so whether visualized as up or down is arbitrary. | (13) Screen image from original LFP simulator. The image in green shows the ideal band locations and the text below it the DNA sizes needed. Not shown are the enzyme choices the program makes for each lane. |
| Planning these images requires knowing what size DNA is required to move at the correct speed in each column of the image. DNA moves through the gelatin at a rate inversely proportional to its size (and moves from the negatively charged end of the gel to the positively charged end of the gel). To determine the proper band-size, a custom simulation programs was written. The LFP simulator first determines what ideal sizes DNA should be to produce the proper bands. This is made possible because many organisms have stable regions of DNA with little variance. Then it exhaustively catalogues the cut points on the DNA that would be made by each possible enzyme and simulates varied possible combinations of these enzymes. The program tries thousands of combinations for each lane of the image and ranks each combination according to each band's deviation from the ideal. Once the best combination is discovered, the program outputs the best combination of enzymes to achieve this. (See figure 1 on "intro" page, showing actual enzymes used for each lane). |
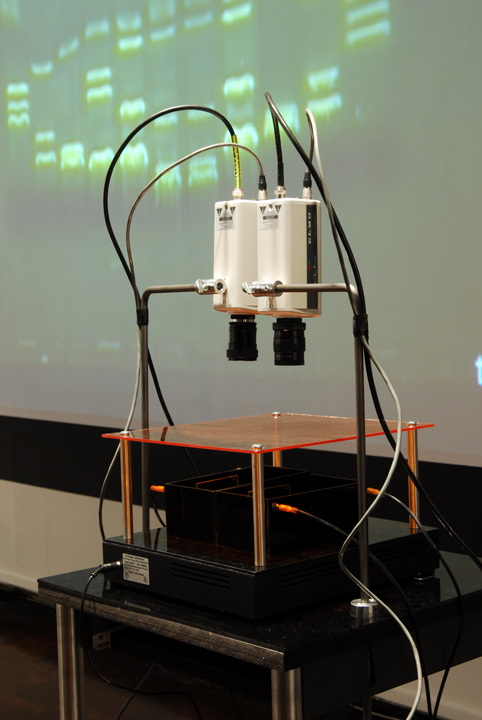
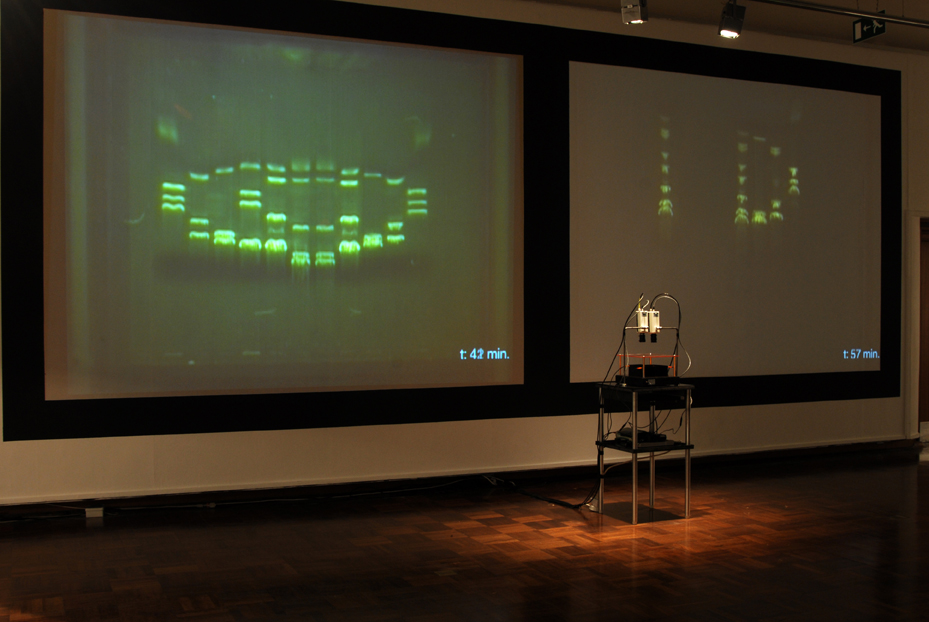

| (14) LFP apparatus and installation, Copywrong, Duncan of Jordanstone College of Art and Design, Dundee, Scotland. | (15) LFP apparatus closeup. |
| The imaging process (gel electrophoresis) is captured live by video cameras attached to the installation worktable. This is achieved through a novel combination of non-hazzardous DNA stain, blue light LEDs and tinted acrylic filters that allow the DNA’s movements to be visible to the naked eye without eye protection, whereas typically in lab-work, the imaging process occurs only at the end of electrophoresis and often requires UV light protection. |